Pandemic puts Oxford Nanopore ‘on the map’

Roula Khalaf, Editor of the FT, selects her favourite stories in this weekly newsletter.
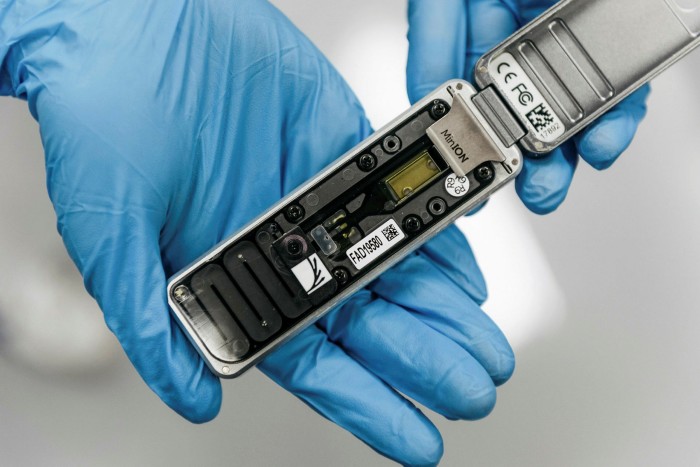
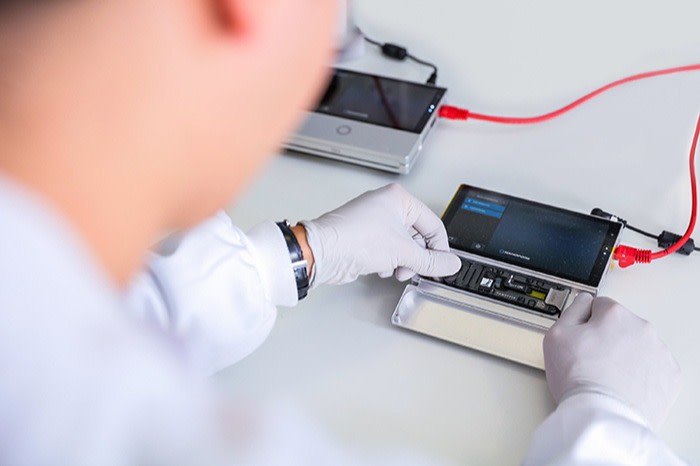
In April 2015, researchers from the University of Birmingham in the UK arrived in Guinea during the Ebola outbreak with nothing but a few suitcases. In one was a mobile phone-sized device that would change the course of the epidemic — a portable DNA sequencer called the MinION.
The device, along with laptops and accessories, enabled the researchers to set up a self-contained laboratory. Instantly, they could examine the evolving virus and its variants, and help shape the development of a novel vaccine.
The sequencer was made by a small UK company called Oxford Nanopore Technologies, which was spun out of the University of Oxford in 2005. In September 2021, in the midst of the Covid-19 pandemic, the company floated on the London Stock Exchange, achieving a market capitalisation of £3.5bn, which made it the third-largest biotechnology listing globally this year.
“Genome-sequencing information is crucial for researchers and epidemiologists during an epidemic,” Nick Loman, the University of Birmingham professor who led the Ebola team, wrote in a report after the epidemic. “Yet, generating such information is a laborious process typically performed in well-equipped laboratories using large, delicate and expensive hardware.”
In Guinea, the Birmingham team discovered that the MinION took less than an hour to sequence a sample of the Ebola virus, meaning they could analyse specimens and produce usable information within 24 hours.
Months later, the team travelled to Brazil to study the spread of the Zika virus using the same sequencers, helping in the public health management of the outbreak. Nanopore sequencing had turned genomic sequencing into a real-time, decentralised community effort that could save lives. So, when the Covid-19 outbreak was declared a pandemic last year, the Oxford-based company’s patented technology proved indispensable. The devices were essential in identifying and tracking the spread of Covid variants in 85 countries, and have sequenced about 18 per cent of all coronavirus genomes globally.
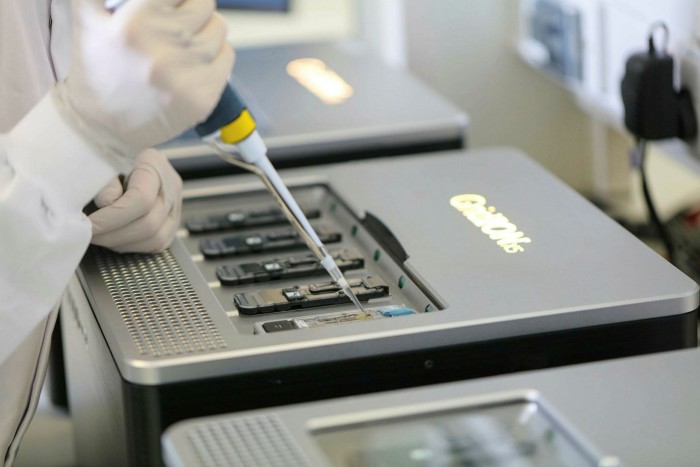

“Oxford Nanopore is a frontrunner in the next generation of sequencing technologies,” says Julian Roberts, an equities analyst at Jefferies. “The last year changed Nanopore’s revenues completely. One contract alone from the UK government was £115m, which was more than the company’s entire revenue for the previous year. The pandemic really put it on the map.”
Oxford Nanopore made a gross profit of £26.9m in the first half of 2021, with a gross margin of 51 per cent, the company says. Revenues rose 22 per cent to £59m in the six months to June, compared with the same period last year.
The $7bn market for DNA sequencing has so far been dominated by Illumina of the US. The company, which has a market cap of $60bn, entered the market by buying a UK start-up called Solexa in 2007. Devices such as Illumina’s can take days to produce sequencing information and are the size of a large filing cabinet and therefore not portable. Nanopore’s devices, which are approaching Illumina’s for accuracy, are cheaper, quicker and have been used everywhere from country hospitals to city streets, farms, and oceans.
“I would have never imagined that a few years after it helped track the Ebola and Zika viruses, we would be doing the same thing in the UK,” says Matt Loose, a developmental biologist at the University of Nottingham who led the Nottingham chapter of the Covid-19 Genomics UK Consortium (COG-UK). “The pandemic was an exceptional event, but we were able to start sequencing Sars-Cov-2 within a couple of weeks of patients appearing in Nottinghamshire hospitals. We took things developed in the field in west Africa and adapted them and scaled them up for use in the UK.”
Oxford Nanopore’s patented technology allows scientists to sequence DNA by creating pores in a synthetic material that are just small enough for a single DNA molecule to be passed through and identified by measuring changes in the electrical current.
During the pandemic, COG-UK teams such as Loose’s used it to analyse the genomes of coronavirus samples in communities and sent the information to local hospitals. They also sent it centrally, to Public Health England and other organisations that were analysing the data at scale, looking at the spread of variants, for instance.

“A group of people met early on and said we need to be running genetic epidemiology of this virus and understanding what is happening globally, and the UK was world-leading,” Loose says. “The impact of that has been that we were able to use nanopore sequencing to follow the variants, see how the virus is changing, and measure and understand its spread.”
Loose’s team of eight scientists received daily samples from patients in Nottinghamshire hospitals and were able to track hospital-acquired transmissions and identify the Delta variant early on, from people who had recently returned from India or Bangladesh.
“We hadn’t even heard about epidemiological surveillance until Covid-19, and Oxford Nanopore was at the forefront of feeding into our understanding of how the virus evolved and where it was becoming more present and dangerous,” says Julia Hawkins, a general partner at early-stage investment firms LocalGlobe and Latitude, which invested in Oxford Nanopore in 2019. “Epidemiological surveillance is here to stay. This is something countries everywhere, working with scientists, will be adopting as a matter of course in future.

“The life sciences tools market is made up of more than 1m life scientists and over 100,000 labs globally. This provides a great foundation for both near- and medium-term growth for Oxford Nanopore.”
The company’s core customers are universities and laboratories conducting scientific research, but it hopes the Covid-19 bump it experienced will help the industry move towards applied genomics, or a “global Internet of Living Things” — real-time networks of biological sensors that can be used in hospitals, pharmaceuticals, food and water safety and agriculture — which it says could be valued in the tens of billions of pounds.
“Covid-19 is changing how broader communities think about infectious disease. With the spectre of microbial resistance approaching, governments are now more aware of the health and economic threat of pathogens, and taking pathogen surveillance much more seriously,” says Gordon Sanghera, Oxford Nanopore chief executive. “The challenge isn’t just understanding pathogens in humans; to have real visibility, epidemiologists would like to build pathogen surveillance in animals and even waste water environments.”

An example of the move from lab to clinic is the UK government-funded project by Genomics England, which is testing whether NHS hospitals can conduct accurate real-time sequencing tests for cancer diagnostics. The project, which uses Oxford Nanopore sequencers, has already been piloted for about 12 months, and Genomics England hopes it will be able to roll it out nationally in the coming year.
“Cancer patients need to have very fast diagnostic turnround times because they have to be treated quickly,” says Parker Moss, chief commercial officer at Genomics England. “This means the sequencers can be in the hospitals, closer to the patients, and we don’t have to motorbike it into a national sequencing lab, which is currently in Cambridge.”
He adds that the nanopore method is also able to read areas of the genome that are difficult to do so with “gold-standard” traditional sequencers such as Illumina’s. So it could identify new areas in the genome for drugs to bind to, giving “new therapeutic options for patients”.
Comments